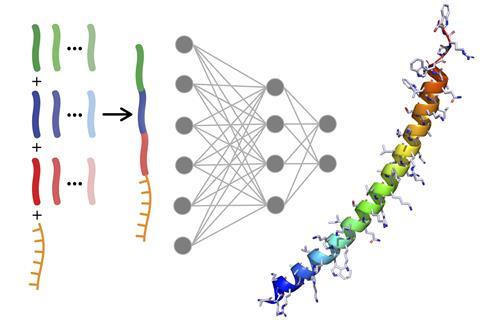
Large biomolecule drugs’ great promise is held back because it’s hard to get them into cells – but now US researchers have used machine learning to tackle this problem. They developed Peptimizer, a machine learning model that designs cell-penetrating peptides (CPPs) that improve oligonucleotide drug delivery into cells 50-fold.
The teams of Rafael Gómez-Bombarelli and Bradley Pentelute at the Massachusetts Institute of Technology worked together with Sarepta Therapeutics, which develops oligonucleotide drugs targeting devastating rare diseases such as Duchenne Muscular Dystrophy (DMD). Oligonucleotides are chains of around 20 nucleotide bases similar to DNA and RNA. This similarity helps oligonucleotides change how genes are turned into proteins, which can benefit genetic disorders like DMD.
While the first oligonucleotide drug was approved in 1998, difficulties getting them into cells have prevented many others reaching the market. This also led to controversy over how well Sarepta’s DMD drug eteplirsen worked when it was approved by the US Food and Drug Administration.
The Pentelute team had found that linking together two known CPPs made from around five to 20 amino acids produced ‘very strong increases in activity’, according to PhD researcher Carly Schissel. However, further improving the CPPs became challenging because there are more combinations of amino acids in a 40-residue sequence than atoms on Earth. The researchers turned to machine learning, showing that a simple neural network could boost oligonucleotide drug delivery threefold.
‘This told us that machine learning could be compatible with what we were doing,’ says Schissel. ‘But we wanted to train a more advanced machine learning model to design really high activity sequences.’
Simply the best training data
Somesh Mohapatra, a PhD researcher in Gómez-Bombarelli’s team therefore developed Peptimizer, a combination of two neural networks. The first was trained with the sequences of over 1000 known CPPs, so that it could suggest new sequences that might be effective.
The researchers fed these sequences to the second, predictor–optimiser neural network. Mohapatra had trained this with data from 600 combinations of oligonucleotide–CPP conjugates that the Pentelute team had synthesised and tested in a cell assay. The consistent conditions offered a clean dataset that’s perfect for machine learning – ‘the best that you can get’, according to Mohapatra. This data taught the neural network to recognise patterns in the sequences that trigger activity. The neural network could predict how active sequences suggested by the generator would be. It could then make small changes and again predict the resulting sequences’ activity.
Pentelute’s team then made 12 CPPs from among the hundreds Peptimizer suggested. While prediction accuracy varied, several new CPPs boosted oligonucleotide delivery more than 20-fold in cell assays, with the best reaching 50-fold. Sarepta researchers also found the CPPs to be effective in mice, where they helped an oligonucleotide reach the animals’ hearts, a key challenge facing existing drugs.
The researchers will now move beyond simple straight CPPs to design different molecule shapes. Mohapatra adds that Peptimizer is applicable to any biological polymer and available as open-source code.
Dominik Heider from the University of Marburg, Germany, calls the work ‘a very good example how AI and machine learning can transform the life sciences and ultimately patient care, drug development, and therapy’. He adds that it’s notable that the team makes useful predictions, and ‘can use their deep learning approach for automated peptide design’.
The study is also a small step towards the chemistry ‘holy grail’ of designing custom enzymes. ‘This is on the path towards deciphering sequence function and possibly structure function,’ says Schissel. ‘But there is still a need for massive amounts of relevant data in order to find completely novel enzymes.’